|
3/22/2023 0 Comments Okazaki fragments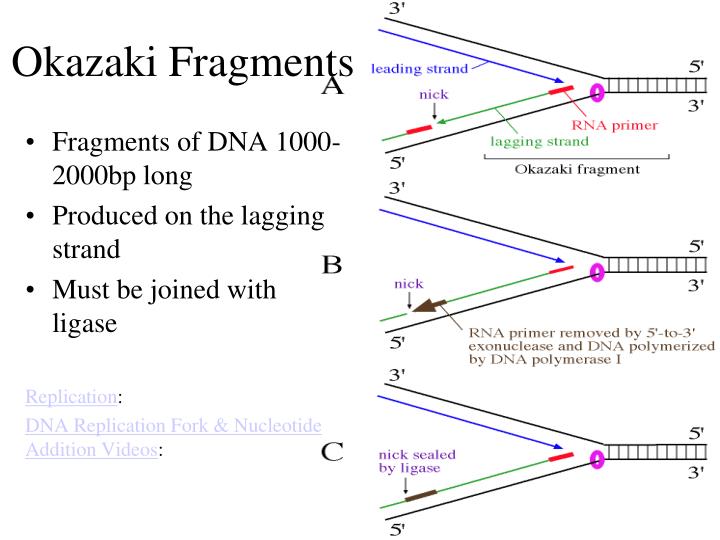
Add reaction mix, chemical, primers, nucleotides and DNA polymeraseģ. It discusses the difference between the leading strand and the lagging stran. While researching the replication of bacteriophage DNA in Escherichia coli in 1968, Reiji Okazaki and his wife, Tsuneko Okazaki, discovered the Okazaki fragments. ligation process where DNA fragments produced using restriction enzymes may be reassembled DNA Polymerase Chain reaction (Steps) 1. This biology video tutorial provides a basic introduction into DNA replication. Heterochromatin tightly coiled DNA, not to be transcribed Euchromatin Less condensed, actively being transcribed. P amino acid exits the E site and A amino acid moves to the P site. Elongation Peptidyl Transferase- Covalently bonds the P amino acid with the A amino acid between the COOH and NH3. translation (genetics) the process whereby genetic information coded in messenger RNA directs the formation of a specific protein at a ribosome in the cytoplasm Initiation In an enzyme-catalyzed reaction, the stage during which enzymes orient reactants precisely as they bind at specific locations within the enzyme's active site. Base pairs w/ mRNA (codon tRNA= anti codon) transcription (genetics) the organic process whereby the DNA sequence in a gene is copied into mRNA. at one end (CCA stem) with amino acid -acyl tRNA synthetaseģ. raw tRNA contains introns which must be excisedġ. These portions are called Okazaki Fragments. undergoes POST-TRANSCRIPTIONAL MODIFICATION On the lagging strand, there are portions of synthesized DNA. (mRNA= 5' to 3') tRNA reads mRNA to be complementary to it. messanger from DNA to cytoplasm to make polypeptides (amino acid sequences). Quito= O Nucleoside Sugar attached to any purine or pyrimidine Nucleotide Phosphate + Sugar + Organic Base Phosphodiester bond refers to the carbon by which the phosphate group is attached rRNA ribosomal RNA type of RNA that makes up part of the ribosome. Trisomy (2n+1) 47 chromosomes Cystine Pyrimidine (can add only) Holandric Traits carried only on the Y chromosome Aneuploidy Cells that have extra chromosomes or chromosomes missing. Comma Splices, Run-ons, and Fragments - UVU. Primer strand of nucleic acid that serves as starting point for DNA synthesis on lagging strand DNA polymerase III adds nucleotide, only in 5' - 3' direction Okazaki fragments are short sequences of DNA nucleotides which are synthesized discontinuously and later linked together by the enzyme DNA ligase to create. Many Okazaki fragments make up the lagging strand of newly synthesized DNA.ġ00-200 nucleotides long in Eukaryotes. 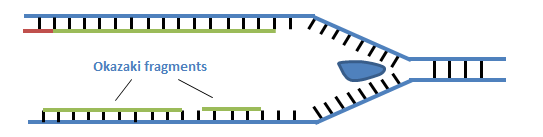
Replication of the leading strand begins near the origin of replication. Replication of the leading strand is continuous. Define the characteristics of the leading and lagging strands in the replication bubble. 
However, Dna2p has a role in a pathway for processing structured flaps, in which it aids FEN1 using both its nuclease and helicase activities.Okazaki fragment A short segment of DNA synthesized on a template strand during DNA replication. Step 3: Define the terms leading strand, lagging strand, Okazaki fragment, and RNA primer. Our results suggest Dna2p is not used for processing of most flaps. The presence of high Dna2p activity, under reaction conditions favoring helicase activity, substantially stimulated FEN1 cleavage of tailed-foldback flaps and also 30-nucleotide unstructured flaps. CTG flaps can form foldback structures and were inhibitory to both nucleases, however, addition of a dT(12) to the 5'-end of a CTG flap allowed Dna2p cleavage. On 30-nucleotide fixed or equilibrating flaps, RPA partially inhibits FEN1. FEN1 cleaves 10-nucleotide fixed or equilibrating flaps in an efficient reaction, insensitive to even high levels of RPA or Dna2p. Reactions required to prime the next fragment, elongate the present fragment, and remove primers from the previous fragments can occur simultaneously. To determine the most likely process, we analyzed cleavage of short and long 5'-flaps. Okazaki fragments are 10003000 bases in length and replication is proceeding at 4001000 bases s 1, so the cycle of reactions on the lagging strand must be completed in a few seconds. RPA dissociates from the resultant short flap, allowing FEN1 cleavage. In the other, the single-stranded binding protein, replication protein A (RPA), coats the flap, inhibits FEN1, but stimulates cleavage by the Dna2p helicase/nuclease. In one model for this process, the flap endonuclease 1 (FEN1) removes the iRNA. Each contains an initiator RNA/DNA primer (iRNA/DNA), which is converted into a 5'-flap and then removed prior to fragment joining. Short DNA segments designated Okazaki fragments are intermediates in eukaryotic DNA replication. On the Roles of Saccharomyces cerevisiae Dna2p and Flap Endonuclease 1 in Okazaki Fragment Processing* Abstract
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
